CDesk: 4. ChIPseq&CUTTag pipeline
Our CDesk ChIPseqCUTTag module comprises of 7 function submodules. Here we present you the CDesk ChIPseqCUTTag working pipeline and how to use it to analyze your CvhIPseq&CUTTag data.
4.1 ChIPseq&CUTTag: Preprocess
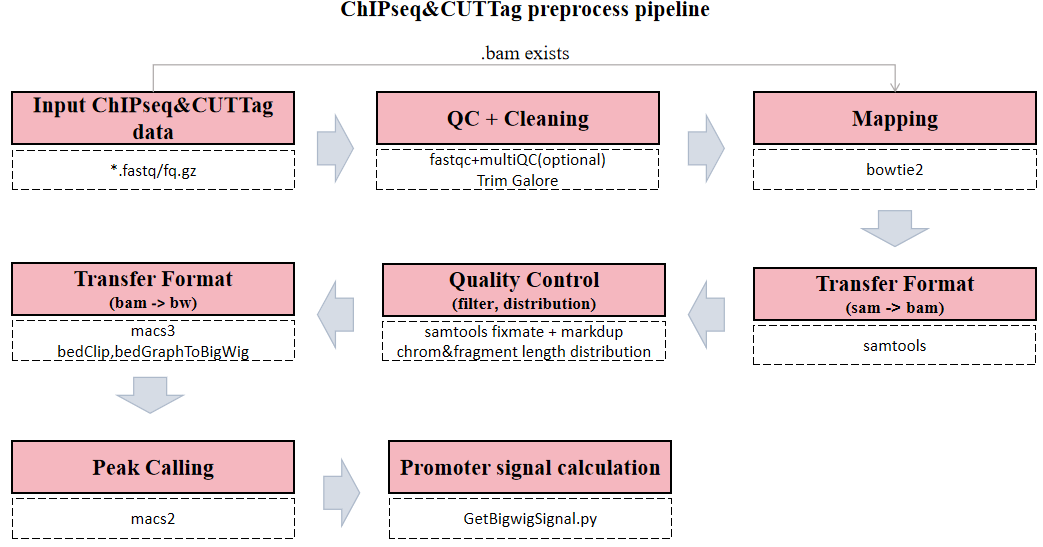
The CDesk ChIPseqCUTTag preprocess pipeline is illustrated in the figure below. The input is a csv file containing metadata of FASTQ files.
The pipeline first checks whether a BAM file corresponding to each FASTQ sequencing file already exists. If so, the alignment step is skipped to save time. Next, each FASTQ file undergoes optional quality control using FastQC and MultiQC to assess key metrics such as base quality distribution, GC content, and adapter contamination. Following quality assessment, Trim Galore is applied to trim low-quality bases and remove adapter sequences, ensuring high accuracy in downstream analyses. Subsequently, Bowtie2 is used to align the cleaned reads to the reference genome, determining the genomic origin of each fragment. The resulting SAM files are then converted into sorted and indexed BAM files using Samtools. After format conversion, reads are filtered to remove low-quality and unaligned reads. MACS3 produces a pileup signal that is converted to a normalized BigWig track after filtering and processing.Fragment length distribution is then analyzed and visualized. Peak calling is performed using MACS2, and finally, signal intensity over promoter regions is computed for downstream interpretation.
Here is an example about how to use the CDesk ChIPseqCUTTag preprocess module.
CDesk ChIPseqCUTTag preprocess \
-i /.../input.csv -o /.../output_directory \
-t 50 -s mm10
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| -i,--input* | The input csv | |
| -o,--output* | The output directory | |
| -s,--species* | The species specified | |
| -t,--thread | The number of threads to use | 8 |
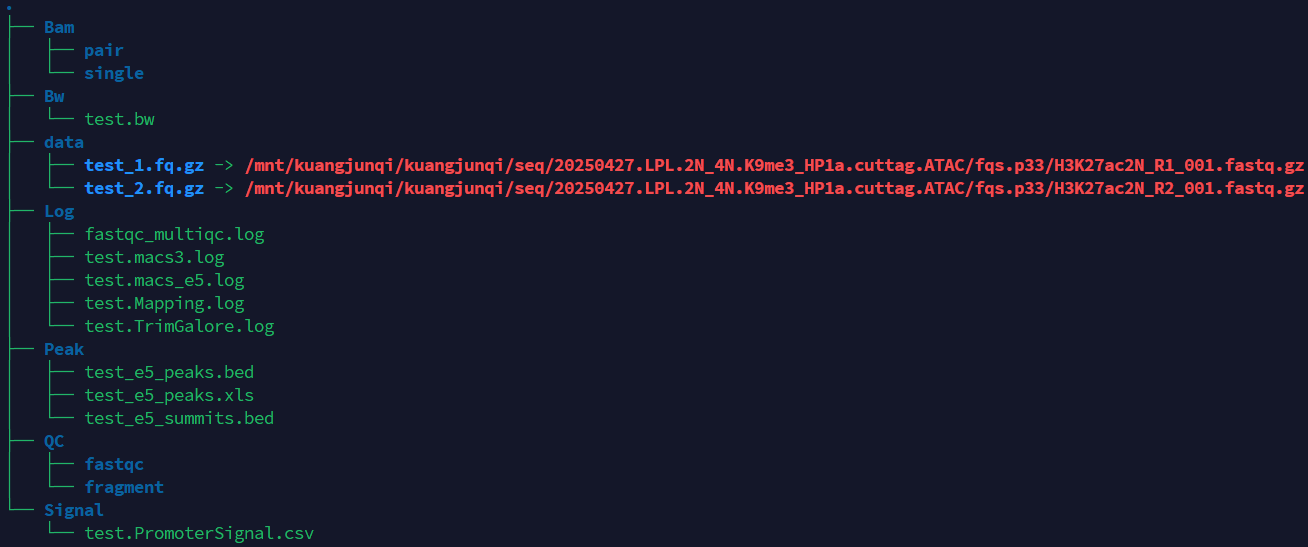
If the pipeline runs successfully, you will see output similar to the figure shown below.
- data: Stores the data path.
- Bam: Stores the intermediate sam and bam files.
- BW: Stores the bw files.
- Log: Stores the log files of fastqc and multiqc, trim_galore, mapping and peak calling.
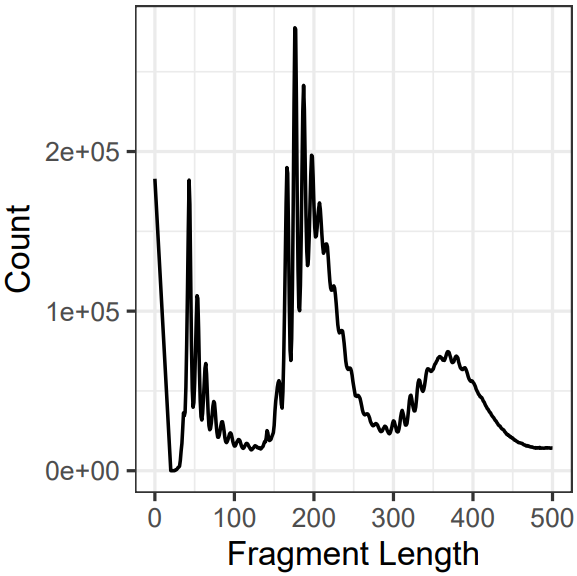
- QC: Stores the fastqc and multiqc result, chromosome distribution and length of fragments.
- Signal: Stores signal strength of promoter regions.
- Peak: Stores the peak files.
A successful CDesk ChIPseqCUTTag preprocess running process
Checking required tools... All required tools are available. --------------------------------------------INITIALIZING---------------------------------------------- ChIPseqCUTTag data analysis pipeline is now running... Number of threads ---------- 100 Directory of data ---------- /mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/new/data Directory of result ---------- /mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/new Mapping index ---------- /mnt/linzejie/data/Drseq_data/bowtie2/mm10/mm10 ChromInfo ---------- /mnt/zhaochengchen/Data/mm10/mm10.len PromoterInfo ---------- /mnt/liudong/data/Genome/mm10/mm10.promoter.ncbiRefSeq.WithUCSC.bed ---------------------------------------Process fq files----------------------------------------------------- ----------------------------------------------------------------------------------------------------- ----------------------------Number 1 fq sample: test-------------------------- ----------------------------------------------------------------------------------------------------- No available BAM file checked,do mapping fastqc 2025-12-08 12:06:17 fastqc ... 1.mapping... paired data 2025-12-08 12:15:26 trim_galore ... paired data 2025-12-08 12:19:54 bowtie2 ... sam2bam 2025-12-08 12:25:40 samtools view ... ---------------------------------------Process bam file---------------------------------------------------- ----------------------------Process test-------------------------- bam2sortbam 2025-12-08 12:26:22 samtools sort ... Sort BAM... [bam_sort_core] merging from 0 files and 100 in-memory blocks... BAM index... Merge replicates... Skip merging bam files ----------------------------Number 1 bam sample: test-------------------------- paired data 2025-12-08 12:33:09 Remove duplicate reads ... [bam_sort_core] merging from 0 files and 100 in-memory blocks... [bam_sort_core] merging from 0 files and 100 in-memory blocks... paired data 2025-12-08 12:44:21 Generate bigwig file ... paired data 2025-12-08 12:52:31 Peak calling ... paired data 2025-12-08 12:58:48 Promoter signal calculation ... Skip 0 transcripts Results saved to /mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/new/Signal/test.PromoterSignal.csv ChIPseqCUTTag preprocessing has been finished!!!
What should the input file look like?
sample,fq1,fq2,bam,ports,group SRR35205586,/mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/SRR35205586/SRR35205586.fastq.gz,,,1, SRR5489315,/mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/SRR5489315/SRR5489315_1.fastq.gz,/mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/SRR5489315/SRR5489315_2.fastq.gz,,2, SRR5489321,/mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/SRR5489321/SRR5489321_1.fastq.gz,/mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/SRR5489321/SRR5489321_2.fastq.gz,/mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/test_result/Bam/pair/SRR5489321.bam,2, 5 columns: - sample: The prefix of generated bam files (x.bam, the prefix of input fq files: x_1.fastq.gz/x_1.fq.gz,x_2.fastq.gz/x_2.fq.gz,x.fastq.gz/x.fq.gz) - fq1: The fastq1 file - fq2: The fastq2 file - ports: The ports of the fastq file (1/2) - bam(optional): The bam files, skip the mapping step if provided - group(optional): The bam files of the same group would be merged if provided
4.2 ChIPseq&CUTTag: Correlation
The CDesk ChIPseqCUTTag correlation module performs correlation analysis on ChIPseq&CUTTag data across different samples to visualize sample relationships. The genome is binned by chromosome at a user-defined bin size. When a peak region file is provided, these regions are merged. Signal similarity across samples is calculated within these regions. PCA and correlation heatmaps are generated to help researchers interpret sample distribution and underlying structure in the data.
Here is an example about how to use the CDesk ChIPseqCUTTag correlation module.
CDesk ChIPseqCUTTag correlation \
-i /.../input.csv -o /.../output_directory \
--bin 100000 -t 100 --species mm10
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| -i,--input* | The input ChIPseqCUTTag sample information file | |
| -o,--output* | The output directory | |
| -s,--species* | The species specified | |
| --bin | Bin size for genomic binning | 100000 |
| --step | Peak file partition step | 1 |
| -t,--thread | The number of threads to use | 30 |
| --width | The plot width | 12 |
| --height | The plot height | 8 |
If the pipeline runs successfully, there would be PCA and correlation heatmap of binned genome or merged peaks.
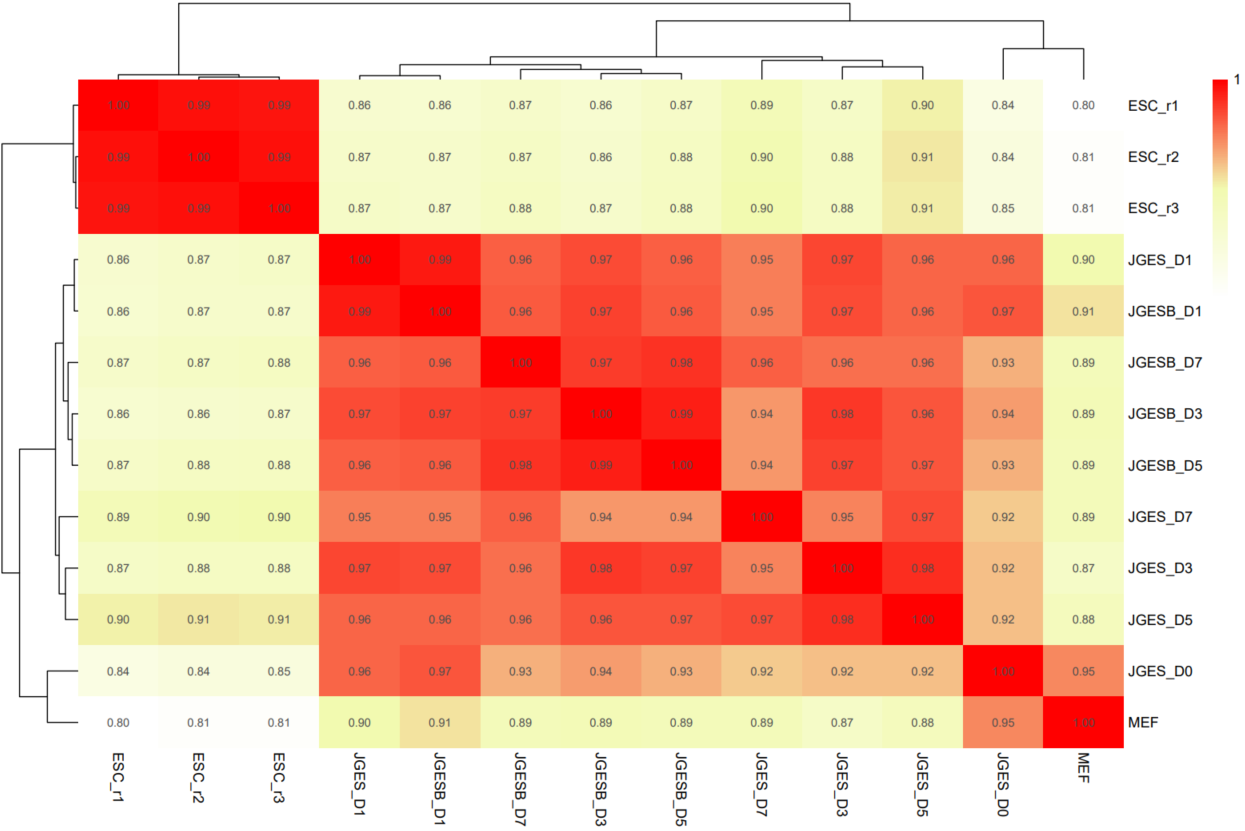
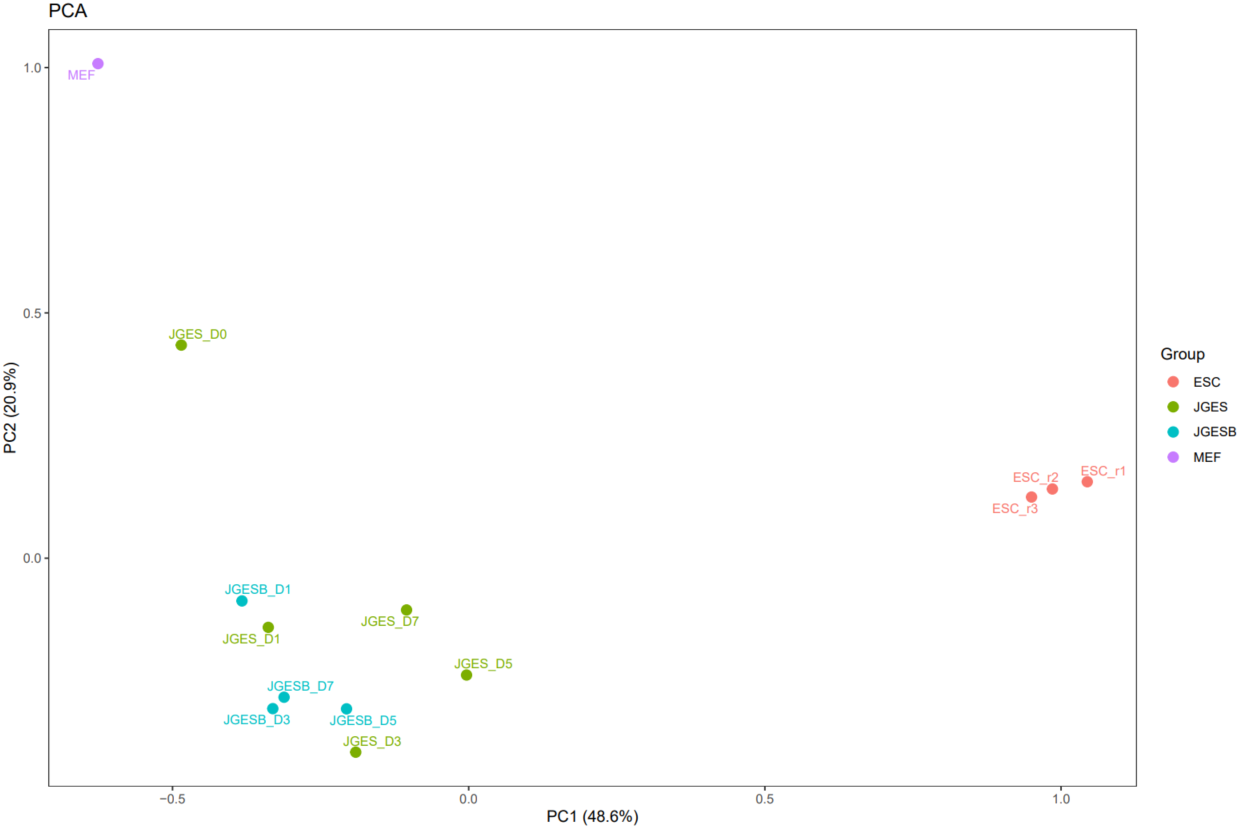
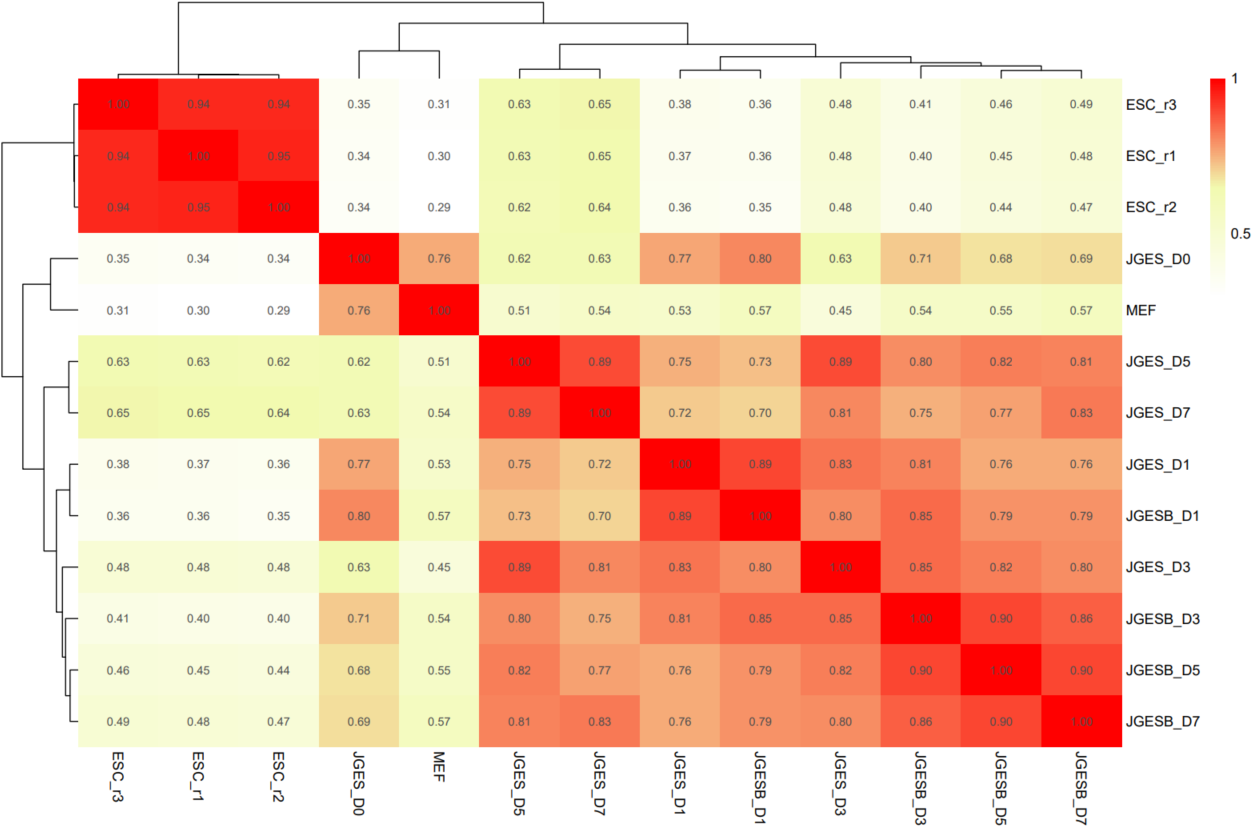
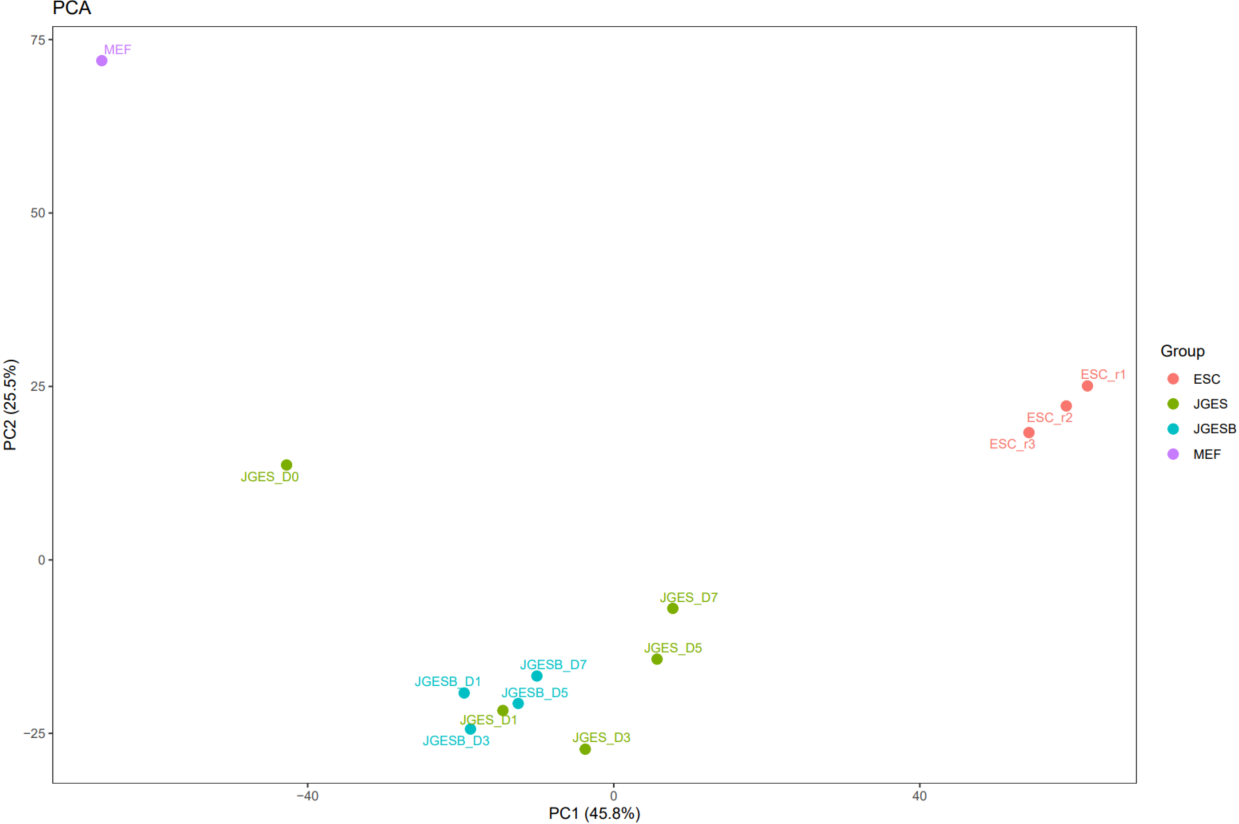
What should the input file look like?
group,tag,bw,peak JGES,JGES_D0,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGES_D0.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGES_D0_peaks.bed JGES,JGES_D1,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGES_D1.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGES_D1_peaks.bed JGES,JGES_D3,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGES_D3.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGES_D3_peaks.bed JGES,JGES_D5,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGES_D5.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGES_D5_peaks.bed JGES,JGES_D7,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGES_D7.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGES_D7_peaks.bed MEF,MEF,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/MEF.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/MEF_peaks.bed JGESB,JGESB_D1,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGESB_D1.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGESB_D1_peaks.bed JGESB,JGESB_D3,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGESB_D3.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGESB_D3_peaks.bed JGESB,JGESB_D5,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGESB_D5.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGESB_D5_peaks.bed JGESB,JGESB_D7,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/JGESB_D7.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/JGESB_D7_peaks.bed ESC,ESC_r1,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/ESC_r1.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/ESC_r1_peaks.bed ESC,ESC_r2,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/ESC_r2.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/ESC_r2_peaks.bed ESC,ESC_r3,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/bw/ESC_r3.bw,/mnt/linzejie/CDesk_test/data/3.ChIPseqCUTTag/2.QC/peak/ESC_r3_peaks.bed - bw: The bw files - group: Same group would be assigned as same color in the PCA plot - tag: Assign the tag in the plots - peak(optional): The peak files to merge if peak parameter assigned
4.3 ChIPseq&CUTTag: Pattern
The CDesk ChIPseqCUTTag pattern module takes as input a list of bigWig signal files and BED region files. Using computeMatrix from deepTools, it calculates a signal intensity matrix over user-defined genomic regions (e.g., promoters, enhancers). Subsequently, plotHeatmap generates a publication-ready PDF heatmap to visualize the distribution patterns of protein binding or epigenetic modifications around these regions. The module supports two analysis modes: reference-point and scale-regions, and allows customization of key parameters, including upstream/downstream extension length, region body length, heatmap height, and number of parallel threads. All outputs—including the matrix file and heatmap—are saved to the specified output directory.
Here is an example about how to use the CDesk ChIPseqCUTTag pattern module.
CDesk ChIPseqCUTTag pattern \
-bw /.../bw.txt -o /.../output_directory \
-bed /.../bed.txt
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| -bw* | The input bw files list file | |
| -bed* | The bed genomic regions to be analyzed list file | |
| -o,--output* | The output directory | |
| --mode | Analysis mode {reference-point,scale-regions} | reference-point |
| --region | The reference point for reference-point {TSS,TES,center} | center |
| -a,--upstream | Length to extend upstream of the genomic region start site (bp) | 2000 |
| -b,--downstream | Length to extend downstream of the genomic region end site (bp) | 2000 |
| -l,--length | Scaled genomic region length (bp) | 3000 |
| --height | Heatmap height | 10 |
| -t,--thread | The number of threads to use | 10 |
If the pipeline runs successfully, there would be a heatmap pdf and a matrix zip file in the output directory.
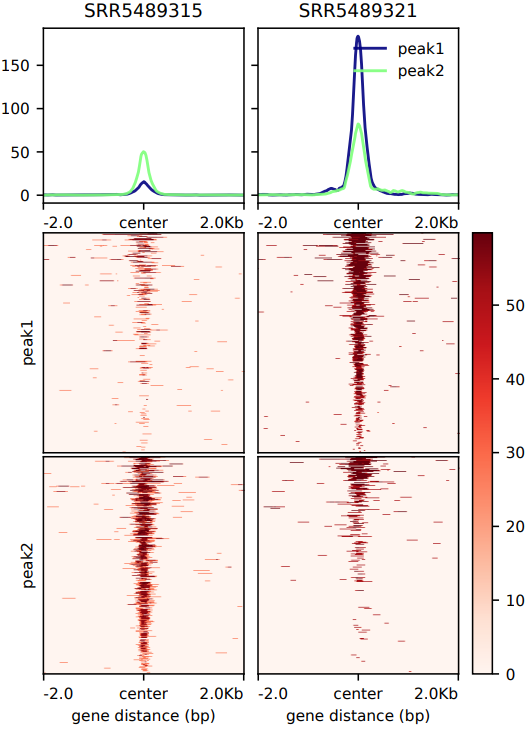
What should the input file look like?
=== bed.txt === /mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/test_result/Peak/peak1 /mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/test_result/Peak/peak2 === bw.txt === /mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/test_result/Bw/SRR5489315.bw /mnt/linzejie/CDesk_test/data/4.ChIPseqCUTTag/1.preprocess/test_result/Bw/SRR5489321.bw
4.4 ChIPseq&CUTTag: Motif
CDesk ChIPseqCUTTag motif module includes four functions for transcription factor motif enrichment analysis:
- HOMER motif discovery – Performs motif analysis on peak regions or gene sequences using the HOMER tool.
Here is an example about how to use the CDesk ChIPseqCUTTag motif homer module.
CDesk ChIPseqCUTTag motif homer \
-i /.../macs_result_peak.csv -o /.../output_directory \
-s mm10 --mode peak
CDesk ChIPseqCUTTag motif homer \
-i /.../gene.txt -o /.../output_directory \
-s mouse --mode gene
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| -i,--input* | The input gene/peak information file | |
| -o,--output* | The output directory | |
| -s,--species* | Available gene in HOMER or provide the path to genome FASTA files for peak mode | |
| --mode* | Analysis mode: peak/gene | |
| -t,--thread | The number of threads to use | 20 |
| --start | offset from TSS (gene mode) | -300 |
| --end | offset from TSS (gene mode) | 50 |
If the pipeline runs successfully, there would be findMotifsGenome, annotatePeaks homer results and log files in the output directory for the peak mode. There would be findMotifs homer results and log files in the output directory for the gene mode.
What should the input file look like?
=== macs_result_peak.csv === peak,sample /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/Find_homer/GSM7789740_e3_peaks.bed,GSM7789740_e3 /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/Find_homer/GSM7789740_e5_peaks.bed,GSM7789740_e5 /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/Find_homer/GSM7789740_e7_peaks.bed,GSM7789740_e7 === gene.csv === gene,sample /mnt/kongtianci/MET/Figure/Supplement/EMT_State_cip_down_genes.txt,EMT_State_cip_down /mnt/kongtianci/MET/Figure/Supplement/EMT_State_cip_up_genes.txt,EMT_State_cip_up - sample: The generated output directory name - peak/gene: The macs call peak results / the genes txt file
- MEME motif analysis – Identifies motifs in FASTA sequences using MEME-ChIP.
Here is an example about how to use the CDesk ChIPseqCUTTag motif meme module.
CDesk ChIPseqCUTTag motif meme \
-i /.../input.csv -o /.../output_directory \
-db /.../db.txt
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| -i,--input* | The input fasta information file | |
| -o,--output* | The output directory | |
| -db,--database* | Database file | |
| -t,--thread | The number of threads to use | 10 |
| --nmotifs | Number of motifs to extract | 3 |
If the pipeline runs successfully, there would be meme results and log files in the output directory.
What should the input file look like?
=== input.csv === fasta,sample /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/Find_meme/GSM7789740_e3_peaks.fa,GSM7789740_e3 /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/Find_meme/GSM7789740_e5_peaks.fa,GSM7789740_e5 /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/Find_meme/GSM7789740_e7_peaks.fa,GSM7789740_e7 - fasta: The fasta files - sample: The sample name=== db.txt === /mnt/kongtianci/CDesk/ACATseq/meme_ref/motif_databases/JASPAR/JASPAR2022_CORE_vertebrates_non-redundant.meme /mnt/kongtianci/CDesk/ACATseq/meme_ref/motif_databases/MOUSE/HOCOMOCOv11_full_MOUSE_mono_meme_format.meme
- HOMER motif discovery result integration and heatmap visualization – Integrates HOMER results to generate TF enrichment heatmaps with significance annotations.
Here is an example about how to use the CDesk ChIPseqCUTTag motif homer_heatmap module.
CDesk ChIPseqCUTTag motif homer_heatmap \
-i /.../input.csv -o /.../output_directory \
--motif /.../motifs.txt
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| -i,--input* | The homer results information csv file | |
| -o,--output* | The output directory | |
| --motif | The specified motifs txt file | no |
| --width | The plot width | 10 |
| --height | The plot height | 8 |
If the pipeline runs successfully, there would a heatmap plot in the output directory. If no motifs specified, it would plot the top 20 significant motifs. (*: q < 0.05, **: q < 0.01, ***: q < 0.001)
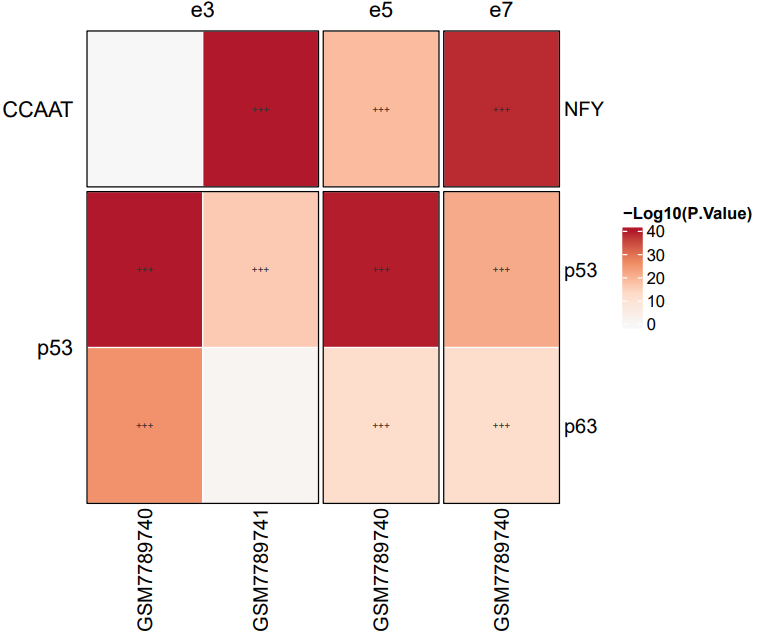
What should the input file look like?
=== input.csv === dir,sample,cluster /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/homer_heatmap/homer_results_test/GSM7789741_e3_peaks_motifDir,GSM7789741,e3 /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/homer_heatmap/homer_results_test/GSM7789740_e5_peaks_motifDir,GSM7789740,e5 /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/homer_heatmap/homer_results_test/GSM7789740_e3_peaks_motifDir,GSM7789740,e3 /mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/homer_heatmap/homer_results_test/GSM7789740_e7_peaks_motifDir,GSM7789740,e7 - dir: homer result directory - sample: The sample name=== motifs.txt === p53 p63 NFY
- Genomic region enrichment analysis – Conducts statistical enrichment testing and visualization of genomic regions associated with motifs.
Here is an example about how to use the CDesk ChIPseqCUTTag enrich_heatmap module.
CDesk ChIPseqCUTTag motif enrich_heatmap \
--motif /.../motifs.csv --bed /.../input.csv\
-o /.../output_directory -t 50 -s mm10
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| --motif_bed* | The homer results information csv file | |
| --bed* | The output directory | |
| -o,--output* | The output directory | |
| -s,--species | The specified species | |
| -t,--thread | The number of threads | 12 |
If the pipeline runs successfully, there would a base directory that saves the input bed files, a MotifAll directory that saves the motif bed files with the statistical enrichment testing result comparing the input and motifs, a summary result file of statistical enrichment testing and the visualization dot plot.
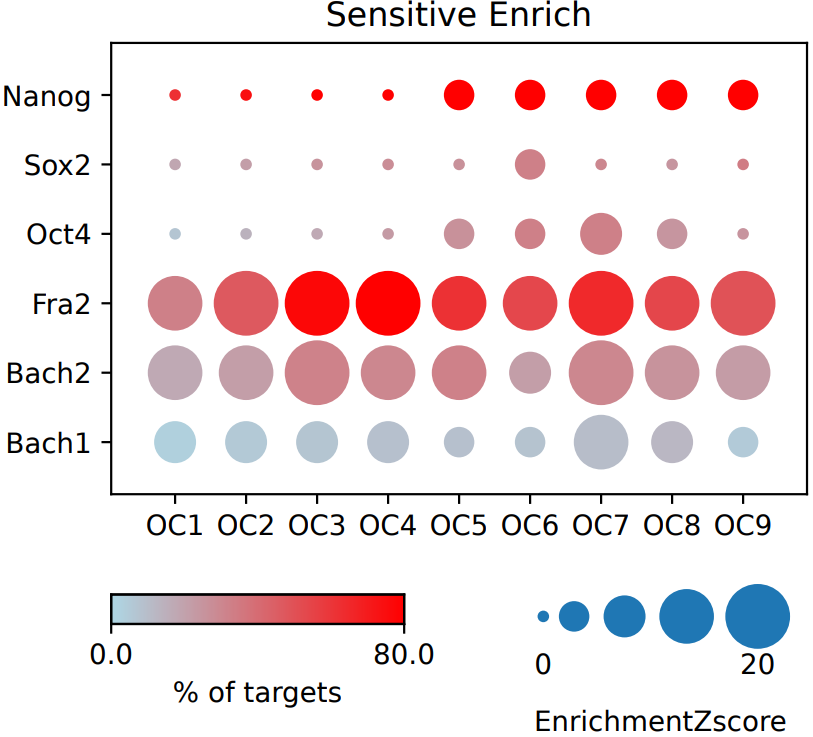

What should the input file look like?
== bed.csv == tag,bed,group OC9,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC9.bed,OC OC8,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC8.bed,OC OC7,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC7.bed,OC OC6,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC6.bed,OC OC5,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC5.bed,OC OC4,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC4.bed,OC OC3,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC3.bed,OC OC2,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC2.bed,OC OC1,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_OC1.bed,OC CO9,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO9.bed,CO CO8,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO8.bed,CO CO7,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO7.bed,CO CO6,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO6.bed,CO CO5,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO5.bed,CO CO4,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO4.bed,CO CO3,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO3.bed,CO CO2,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO2.bed,CO CO1,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/bed/Oct4NanogN70_CO1.bed,CO - tag: The tag to plot - bed: The input bed files - group: The group to separate plots== motif.csv == motif,bed Bach1,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/motif/Bach1.bed Bach2,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/motif/Bach2.bed Sox2,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/motif/Sox2.bed Nanog,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/motif/Nanog.bed Oct4,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/motif/Oct4.bed Fra2,/mnt/linzejie/CDesk_test/data/3.ATAC/4.motif/enrich_heatmap/motif/Fra2.bed - motif: The motifs tag to plot - bed: The input motif bed files
4.5 ChIPseq&CUTTag: Accessbility
The CDesk ChIPseqCUTTag accessbility module is designed to extract signal values from BigWig files and generate average signal profile plots and boxplots. The script supports two analysis modes: region mode and gene mode. In gene mode, species information is required to retrieve gene annotations (GTF file), and the promoter region size can be user-defined. The script computes signal intensity based on user-specified alignment anchors (e.g., TSS, TTS, or peak center), generates average signal profiles, and calculates mean signal values for each region in each sample. Finally, boxplots are generated according to the user’s choice of grouping method—either by sample or by region list—enabling flexible visualization of chromatin accessibility patterns.
Here is an example about how to use the CDesk ChIPseqCUTTag accessbility module.
CDesk ChIPseqCUTTag accessbility \
--bw /.../bw.txt -o /.../output_directory \
-i /.../gene_list.txt \
--mode gene --species mm10 --center whole(TSS/TTS) \
(--method region_list --region_list /.../region_list.txt)
CDesk ChIPseqCUTTag accessbility \
--bw /.../bw.txt -o /.../output_directory \
-i /.../regions.bed \
--mode region --center position(peak) \
(--method region_list --region_list /.../region_list.txt)
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| --bw* | The input bw txt file | |
| -o,--output* | The output directory | |
| --mode* | Analysis mode: region/gene | |
| -i,--input* | The Input gene region or gene list txt file | |
| -s,--species | The species specified (necessarry in gene mode) | |
| --center* | Average profile alignment:region mode:peak/position,gene mode:TSS/TTS/whole | |
| --method | Boxplot plotting method:sample/region_list | sample |
| --promoter_size | Gene promoter region size, default: TSS upstream and downstream 5kb | 5000 |
| --region_list | Region list file for box plot |
If the pipeline runs successfully, there would a box plot, a summary file of signals on different regions, a directory that saves the profile plot and signal matrix.
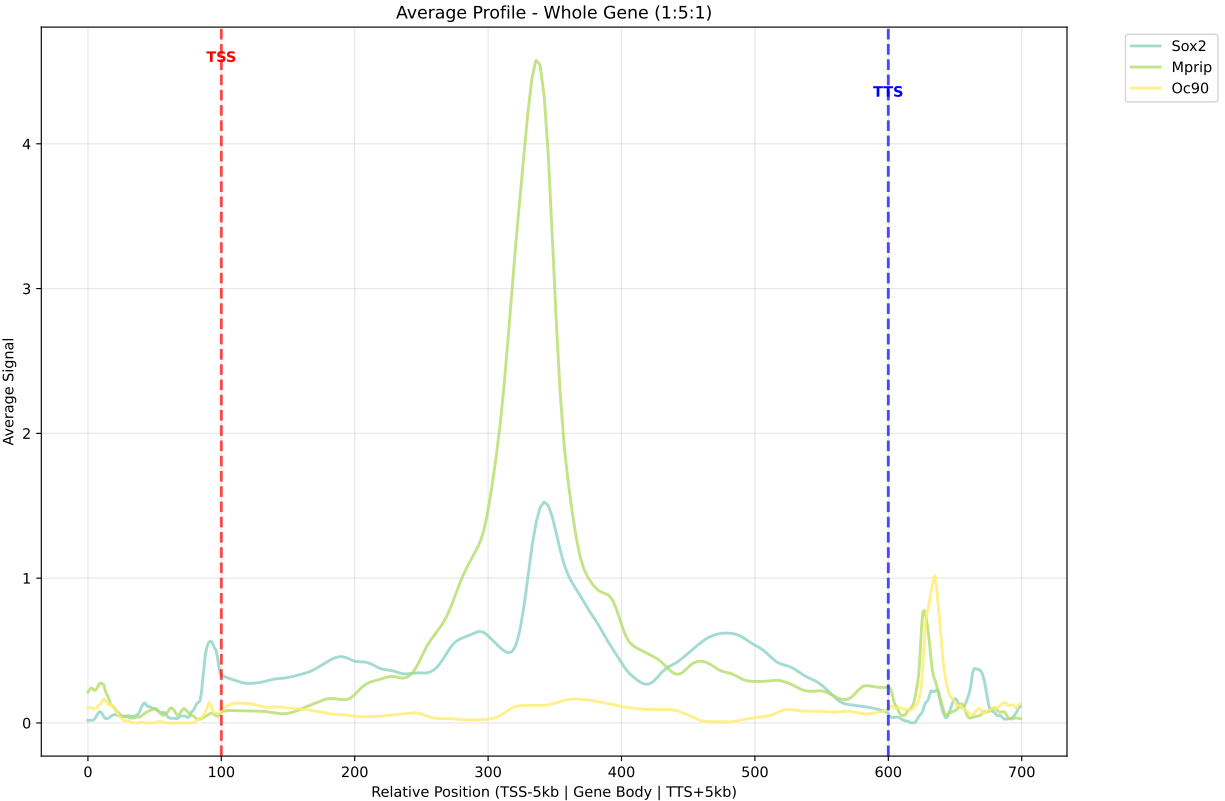
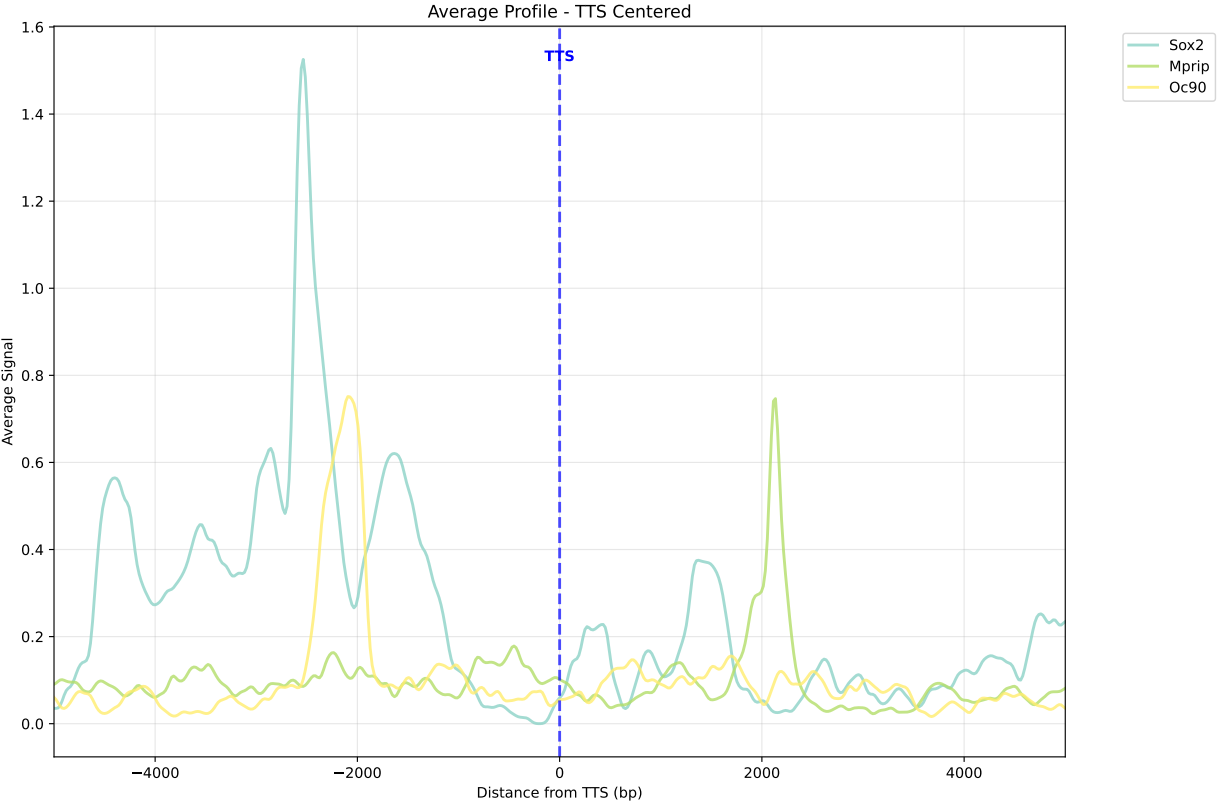
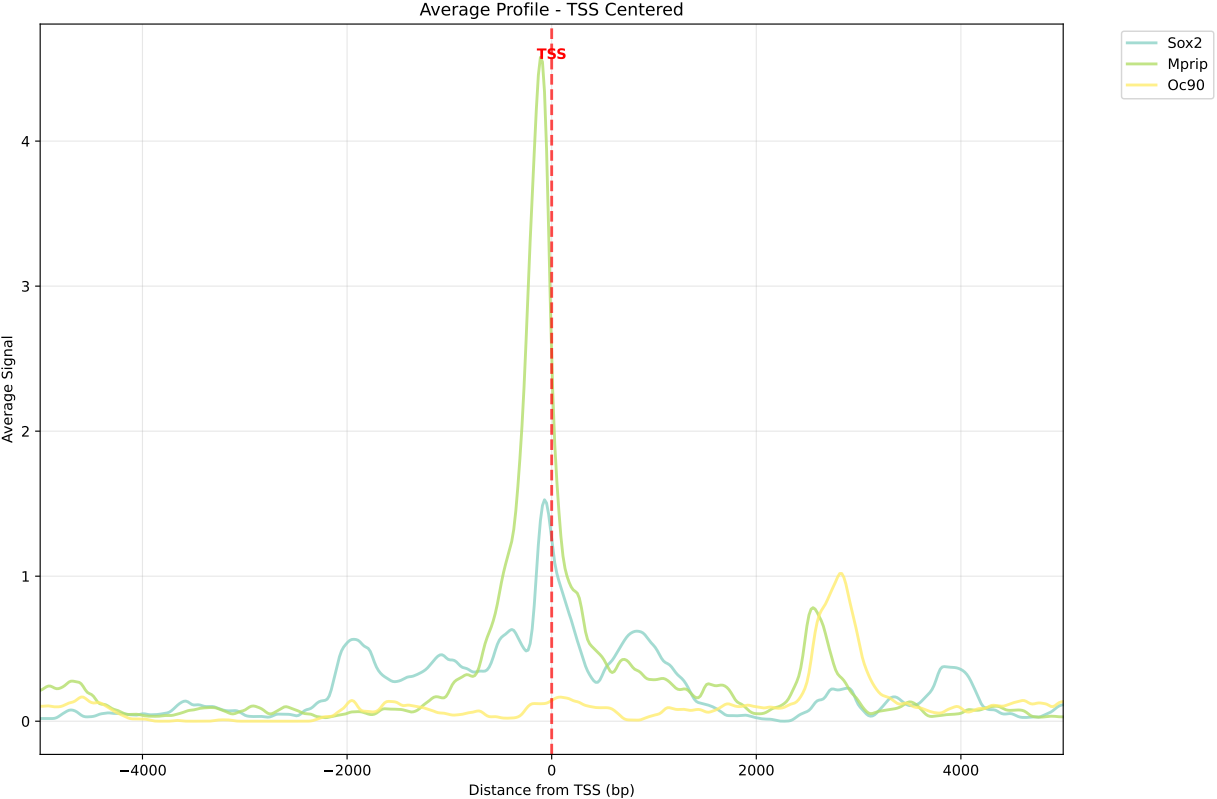
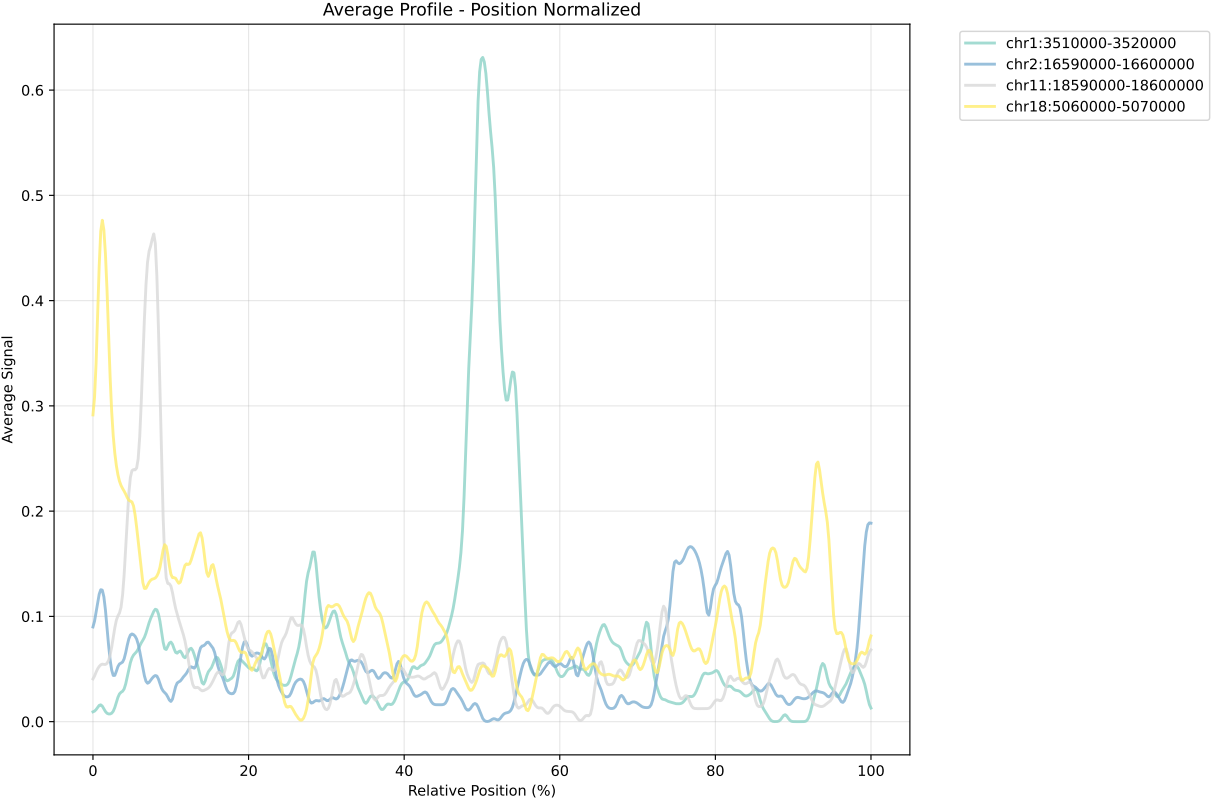
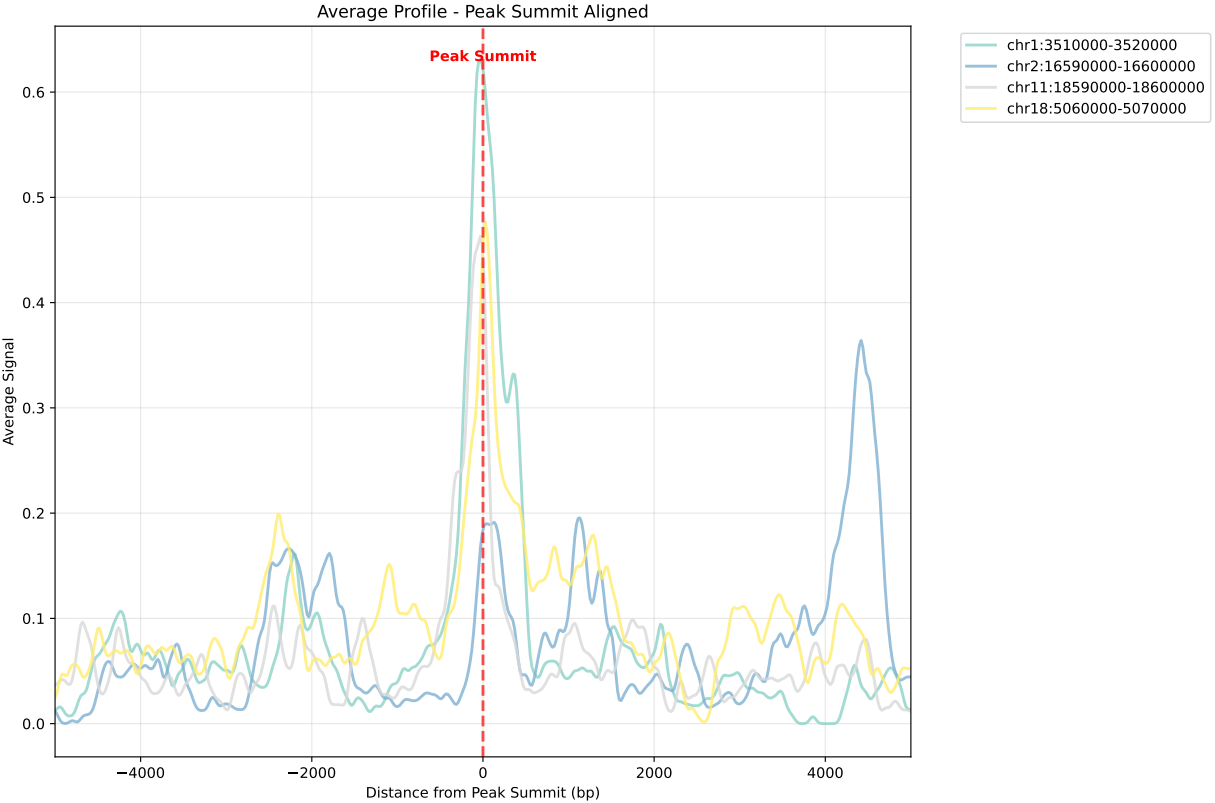
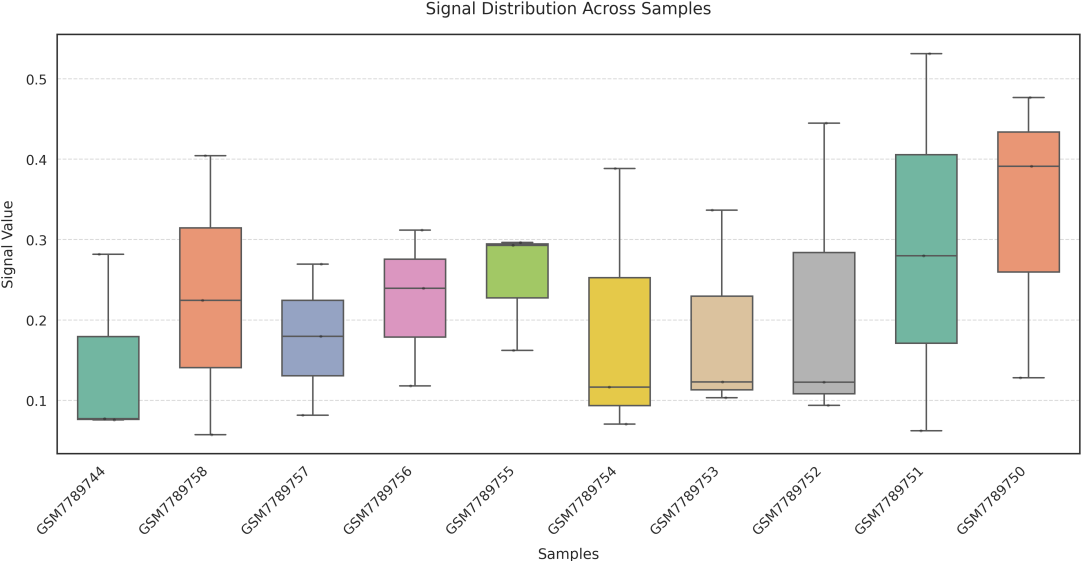
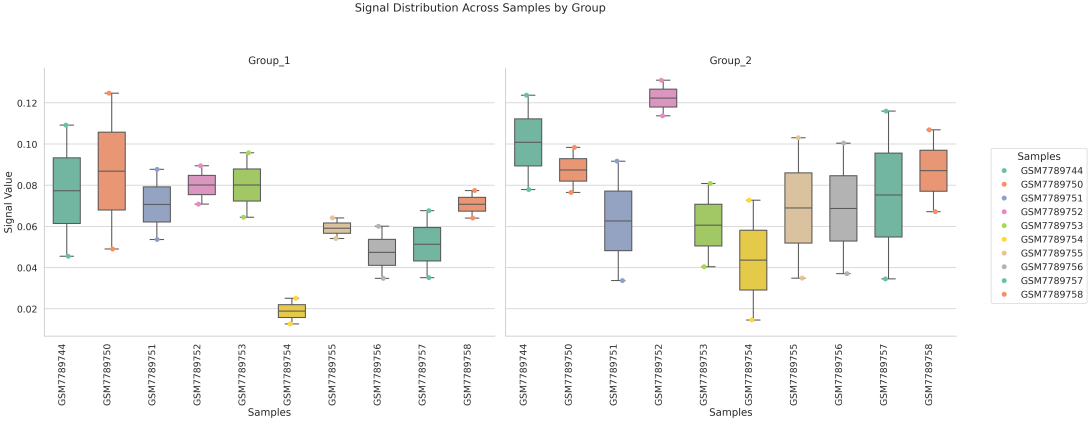
In gene mode, the profile output plot supports centering options: TSS, TTS and whole.
For TSS and TTS, the plot displays the region spanning ±5 kbp around the TSS, TTS, respectively.
For the whole center, the plot includes the entire gene body plus 5 kbp upstream and downstream. Since gene body lengths vary, the gene body region is scaled to a uniform length during visualization to ensure alignment across genes.
Two methods are available for generating the boxplot:
sample: computes the average signal across all genes or regions for each sample, and plots one box per sample;
region_list: groups regions based on a user-provided --region_list file and calculates average signal values for each group, generating a boxplot where each box represents a defined region set.
What should the input file look like?
== bw.txt == /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789744.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789758.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789757.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789756.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789755.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789754.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789753.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789752.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789751.bw /mnt/linzejie/CDesk_test/data/3.ATAC/5.accessbility/GSM7789750.bw == gene_list.txt == Sox2 Mprip Oc90 == region_list.txt ==(each line a group) chr1:3510000-3520000 chr2:16590000-16600000 chr11:18590000-18600000 chr18:5060000-5070000 == region_list.txt ==(each line a group) Sox2 Oc90 Mprip
4.6 ChIPseq&CUTTag: IGV
The CDesk ChIPseqCUTTag igv module is designed to automate the generation of genome browser screenshots using IGV (Integrative Genomics Viewer). It parses a list of input files and specified genomic regions to generate an IGV batch script. The script then runs IGV in a virtual display environment, enabling headless operation, and automatically captures screenshots of the specified genomic regions in bulk.
Here is an example about how to use the CDesk ChIPseqCUTTag igv module.
CDesk ChIPseqCUTTag igv \
-i /.../input.txt -o /.../output_directory \
--region /.../regions.txt --genome hg19
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| -i,--input* | The input list file | |
| -o,--output* | The output directory | |
| --genome* | The genome specified | |
| --region* | Region list file | |
| --display_mode | Display mode:collapse/expand/squish | collapse |
| --type | Output plot format: svg/png | png |
If the pipeline runs successfully, there would an IGV batch script and IGV snapshots in the output directory.
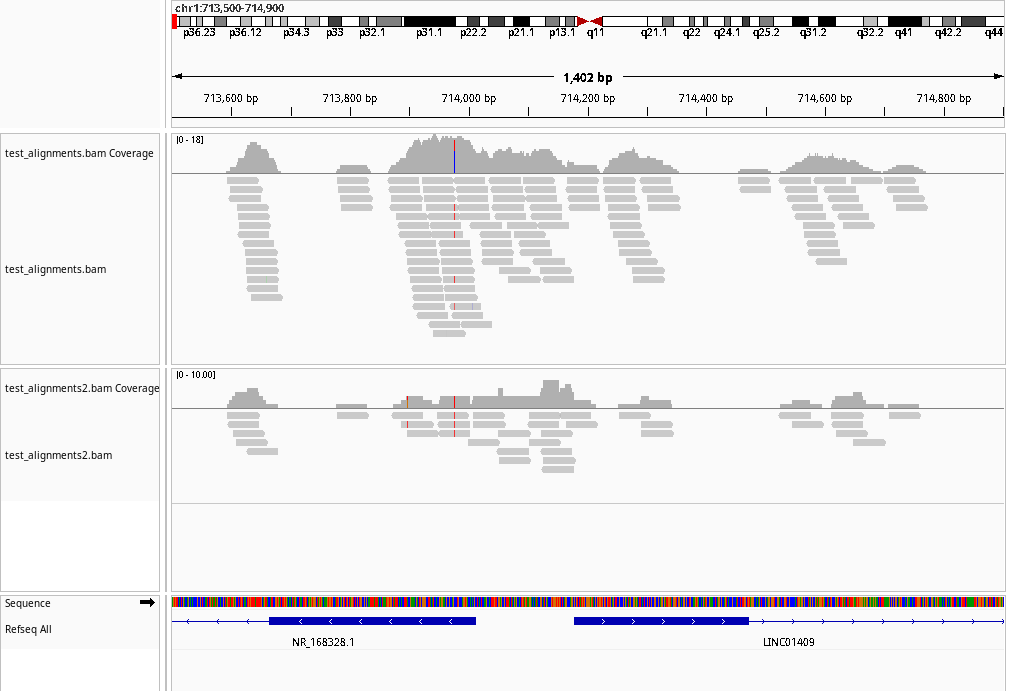
A successful CDesk ChIPseqCUTTag preprocess running process
Using system JDK. IGV requires Java 21. openjdk version "21.0.6" 2025-01-21 OpenJDK Runtime Environment JBR-21.0.6+9-895.97-nomod (build 21.0.6+9-b895.97) OpenJDK 64-Bit Server VM JBR-21.0.6+9-895.97-nomod (build 21.0.6+9-b895.97, mixed mode, sharing) WARNING: Unknown module: jide.common specified to --add-exports WARNING: Unknown module: jide.common specified to --add-exports WARNING: Unknown module: jide.common specified to --add-exports WARNING: Unknown module: jide.common specified to --add-exports WARNING: Unknown module: jide.common specified to --add-exports WARNING: package com.sun.java.swing.plaf.windows not in java.desktop WARNING: package sun.awt.windows not in java.desktop WARNING: Unknown module: jide.common specified to --add-exports WARNING: Unknown module: jide.common specified to --add-exports INFO [Oct 24,2025 10:59] [Main] Startup IGV Version user not_set INFO [Oct 24,2025 10:59] [Main] Java 21.0.6 (build 21.0.6+9-b895.97) 2025-01-21 INFO [Oct 24,2025 10:59] [Main] Java Vendor: JetBrains s.r.o. https://openjdk.org/ INFO [Oct 24,2025 10:59] [Main] JVM: OpenJDK 64-Bit Server VM JBR-21.0.6+9-895.97-nomod INFO [Oct 24,2025 10:59] [Main] OS: Linux 5.15.0-139-generic amd64 INFO [Oct 24,2025 10:59] [Main] IGV Directory: /mnt/linzejie/igv INFO [Oct 24,2025 10:59] [OAuthUtils] Loading Google oAuth properties INFO [Oct 24,2025 10:59] [CommandListener] Listening on port 60151 SLF4J: Failed to load class "org.slf4j.impl.StaticLoggerBinder". SLF4J: Defaulting to no-operation (NOP) logger implementation SLF4J: See http://www.slf4j.org/codes.html#StaticLoggerBinder for further details. INFO [Oct 24,2025 10:59] [BatchRunner] Executing batch script: /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/igv_batch_script.txt INFO [Oct 24,2025 10:59] [GenomeManager] Loading genome: https://raw.githubusercontent.com/igvteam/igv-genomes/refs/heads/main/json/hg38.json SEVERE [Oct 24,2025 11:00] [ReferenceFrame] Null chromosome: INFO [Oct 24,2025 11:00] [TrackLoader] Loading resource: https://hgdownload.soe.ucsc.edu/goldenPath/hg38/database/ncbiRefSeqSelect.txt.gz INFO [Oct 24,2025 11:01] [TrackLoader] Loading resource: https://hgdownload.soe.ucsc.edu/goldenPath/hg38/database/ncbiRefSeqSelect.txt.gz INFO [Oct 24,2025 11:03] [GenomeManager] Loading genome: https://raw.githubusercontent.com/igvteam/igv-data/refs/heads/main/genomes/legacy/json/hg19.json INFO [Oct 24,2025 11:03] [TrackLoader] Loading resource: https://hgdownload.soe.ucsc.edu/goldenPath/hg19/database/ncbiRefSeq.txt.gz INFO [Oct 24,2025 11:05] [TrackLoader] Loading resource: /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam WARNING: BAM index file /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam.bai is older than BAM /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam WARNING: BAM index file /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam.bai is older than BAM /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam WARNING: BAM index file /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam.bai is older than BAM /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam INFO [Oct 24,2025 11:05] [TrackLoader] Loading resource: /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments2.bam Snapshot generation completed successfully.
What should the input file look like?
== input.txt == /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments2.bam == region.txt == chr1:713500-714900 chr3:82500-84000
4.7 ChIPseq&CUTTag: Similarity
The CDesk ChIPseqCUTTag similarity module analyzes the similarity between 2 ATAC-seq samples by computing the genome-wide coverage correlation of two bigWig files using the multiBigwigSummary tool. It supports two modes: genome-wide binning or user-defined region-based analysis. The module calculates both Spearman and Pearson correlation coefficients, filters out regions with zero signal, and generates scatter plots for visual inspection of the correlations.
Here is an example about how to use the CDesk ChIPseqCUTTag similarity module.
CDesk ChIPseqCUTTag similarity \
--bw1 /.../sample1.bw --bw2 /.../sample2.bw \
-o /.../output_directory --region /.../region.bed
| Parameters(*necessary) | Description | Default value |
|---|---|---|
| --bw1* | The first bw file | |
| --bw2* | The second bw file | |
| -o,--output* | The output directory | |
| -t,--thread | Numer of threads | 30 |
| --region | Optional BED file for specific regions |
If the pipeline runs successfully, there would be multiBigwigSummary result files and scatter plots in the output directory.
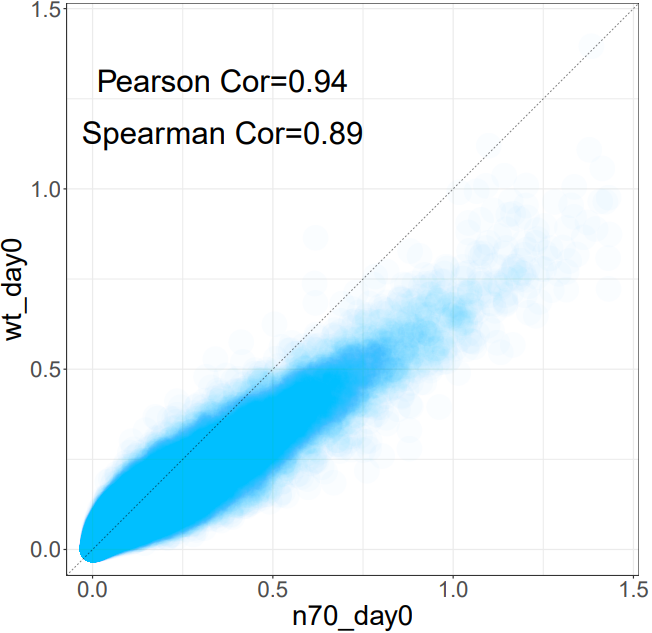
What should the input file look like?
== input.txt == /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments.bam /mnt/linzejie/CDesk_test/data/3.ATAC/6.igv/test_alignments2.bam == region.txt == chr1:713500-714900 chr3:82500-84000